Gene Interaction Network
Biomedical-Bioinformatics, a division of CD Genomics, provides customers with high-quality residue interaction network analysis, which aims to help researchers or doctors systematically explore gene functions in disease research and drug research.
Introduction
With the rapid development of high-throughput biological experiment technology, especially the development of gene chips and next-generation sequencing technology, gene expression data in the whole genome has exploded, and the method of network biology is used to analyze high-throughput gene expression data. Analysis and mining have become an important research direction of bioinformatics. Gene co-expression is a type of analysis method that uses a large amount of gene expression data to construct correlations between genes, thereby mining gene functions. Gene co-expression network is a means to show the interaction relationship between genes, which is based on the expression data between genes to construct a regulatory network diagram. Through gene co-expression network analysis method, functionally related genes can be identified as a module. Through further analysis of the module, advanced analysis such as screening of the core genes of the module, associated traits, metabolic pathway modeling, and the establishment of gene interaction networks can be achieved.
Application Fields
In the field of biomedicine, the gene co-expression interaction network analysis is mainly used in the occurrence and development of diseases research:
- Through the similarity of gene expression, the possible interaction relationships of gene products can be analyzed, so as to understand the interaction context between genes and find core genes.
- Gene function annotation. In biology, genes that exist in the same pathway will show co-expression patterns in terms of expression values, and this feature allows gene function annotations.
- According to the function of genes, key node genes are related to the occurrence and development of diseases.
What We Offer
As one of the global bioinformatics data analysis service providers, CD Genomics provides established, cost-efficient, and rapid turnaround analysis services for gene interaction network analysis for doctors and researchers. CD Genomics provides different software or tools such as WGCNA, Coexpedia, Cytoscape, and other cutting-edge software to perform gene interaction network analysis and visual analysis for researchers to meet the personalized needs. In addition, you only need to provide us with raw data or data analysis requirements. We will be responsible for all follow-up matters of the project, and finally provide you with a complete and easy-to-understand analysis result report. We will generate high-quality charts summarzing the results that can be directly used for article publishing.
Data Analysis Technical Route
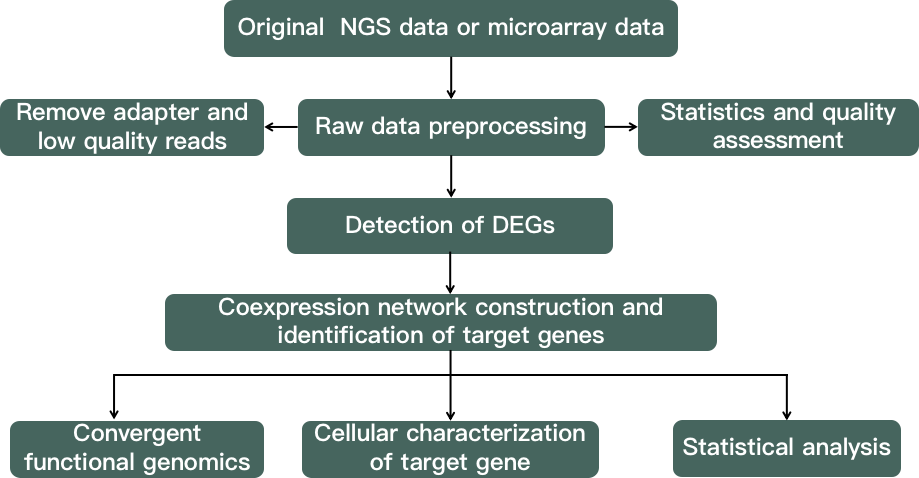 Fig 2. A flow chart demonstrating the gene co-expression interaction network analysis.
Fig 2. A flow chart demonstrating the gene co-expression interaction network analysis.
Service Process
Before data analysis, the first thing is to get your data ready. For gene interaction network analysis, the raw input data can be next-generation sequencing data or microarray data. We will formulate a suitable analysis plan based on your data situation, our service process is as follows:
-
Upload NGS Data or Microarray data
-
Develop an analysis plan
-
Gene co-expression interaction network analysis
-
Generate charts
-
Report interpretation service
For gene interaction network analysis services, if you have any questions about the data analysis cycle, analysis content and price, please click online inquiry.
What's More
For gene interaction network analysis, if you don't have the raw data, CD Genomics relies on years of sequencing experience to provide you with sequencing services, including next-generation sequencing and microarray services. In addition, we also provide services to obtain raw data from public databases. In short, we are happy to work with you at every stage of your research to ensure the best outcome for your study. For gene interaction network, if you have any questions, please feel free to contact us for details.
References
- Xu M, et al. A systematic integrated analysis of brain expression profiles reveals YAP1 and other prioritized hub genes as important upstream regulators inAlzheimer's disease[J]. Alzheimers & Dementia, 2017. 1-14.
- Feng C, et al. Genes related to the very early stage of ConA-induced fulminant hepatitis: a gene-chip-based study in a mouse model[J]. Bmc Genomics, 2010, 11(1):240.
* For research use only. Not for use in clinical diagnosis or treatment of humans or animals.
Online Inquiry
Please submit a detailed description of your project. Our industry-leading scientists will review the information provided as soon as possible. You can also send emails directly to for inquiries.
